A research team led by Sun Yat-sen University Cancer Center (SYSUCC) has published a landmark study in Nature on April 15, 2026, titled "EBV strain interacts with host HLA to drive nasopharyngeal carcinoma risk." The research cracks a long-standing mystery: why is nasopharyngeal carcinoma (NPC) so prevalent in Southern China despite Epstein-Barr virus (EBV) infecting over 95% of adults worldwide? The answer lies not in a single factor, but in a specific genetic interaction between a high-risk EBV variant and host HLA-A alleles.

The study was co-led by Prof. Miao Xu from SYSUCC, along with Prof. Jianjun Liu (Genome Institute of Singapore), Prof. Weiwei Zhai (Institute of Zoology, Chinese Academy of Sciences), and Prof. Zhonghua Liu (Columbia University).
The Puzzle: Why Does NPC Favor Southern China?Although EBV infects the vast majority of the global population, NPC remains endemic specifically in Southern China—an area home to only 2% of the world's population yet bearing over 40% of global NPC cases. For decades, researchers debated whether this "geographic curse" was due to host genetic susceptibility or a locally prevalent high-risk EBV subtype.
The Breakthrough: Identifying a Key Host-Virus InteractionThrough a stepwise host–EBV genome interaction analysis, the team identified a genetic interaction between HLA-A*11:01 and the high-risk EBV variant 85841G as a key determinant of NPC risk.
Individuals carrying a susceptible HLA-A background (HLA-A*11:01− or HLA-A*02:07+) and infected with the high-risk 85841G EBV form a dual-risk subgroup with substantially elevated, interaction-driven NPC risk, far exceeding the effects of host or virus alone. This dual-risk subgroup comprises 20.5% of the population and accounts for approximately 47% of NPC cases.
Mechanism: How the Interaction Drives RiskMechanistically, the EBV 85841G variant encodes an EBNA3B peptide that binds to HLA-A*11:01 and elicits specific T cell responses capable of lysing EBV+ B cells transformed by 85841G-carrying strains. Notably, this interaction is associated with reduced salivary viral load and lower NPC risk among A*11:01 carriers. In other words, the HLA-A*11:01 allele helps the immune system recognize and control the high-risk virus, while its absence leaves carriers vulnerable.
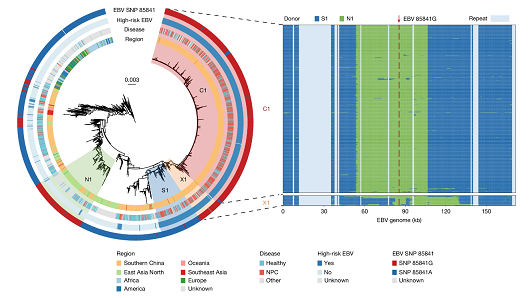
Evolutionary Origin: Why the High-Risk Variant Thrives in Southern ChinaEvolutionary analysis reveals that 85841G arose via ancient recombination events between northern and southern EBV lineages and subsequently underwent clonal expansion in Southern China. This led to the co-enrichment of interacting host and viral risk factors that, together, contribute to NPC endemicity in the region.
Implications: Toward Precision Prevention
This research transforms our understanding of NPC etiology by revealing a markedly stratified, interaction-driven risk architecture. By identifying precisely which host-virus combinations confer the greatest risk—a 17-fold increase compared to baseline—the findings provide a clear roadmap for future precision prevention strategies. These include accurate risk stratification, identification of high-risk populations for regular surveillance, and the development of effective EBV vaccines targeting the high-risk variant.
Authorship and FundingCorresponding authors: Prof. Miao Xu (SYSUCC), Prof. Jianjun Liu (Genome Institute of Singapore), Prof. Weiwei Zhai (Institute of Zoology, Chinese Academy of Sciences), and Prof. Zhonghua Liu (Columbia University).
Co-first authors: Dr. Yanhong Chen and Dr. Jingtong Liang (SYSUCC).
Original link to the article:
https://www.nature.com/articles/s41586-026-10416-8?sessionid=-1466574011

